|
Mirex$year 0.2,1,0)įSA::peek(Mirex,n=10) # examine a portion of the data frame # year weight mirex species gt2 These same data were used in this post about depredating fitPlot(). 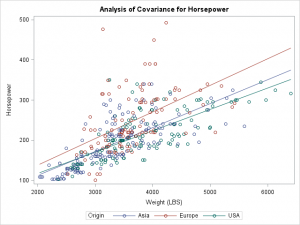
The year variable is converted to a factor below for modeling purposes. Most examples below use the Mirex data set from FSA, which contains the concentration of mirex in the tissue and the body weight of two species of salmon ( chinook and coho) captured in six years. Library(patchwork) # placing plots (in conclusion) Library(FSA) # fitPlot() code may not run after >v0.9.0 library(tidyverse) # for dplyr and ggplot2 The examples below require the following additional packages. The basic plots produced by residPlot() are recreated here using ggplot2 to provide a resource to help users that relied on residPlot() transition to ggplot2. We now feel that students are better served by learning how to create these visualizations using methods provided by ggplot2, which require more code, but are more modern, flexible, and transparent. residPlot() was originally designed for students to quickly visualize residuals from one- and two-way ANOVAs and simple, indicator variable, and logistic regressions. We are taking this action to make FSA more focused on fisheries applications and to eliminate “black box” functions. 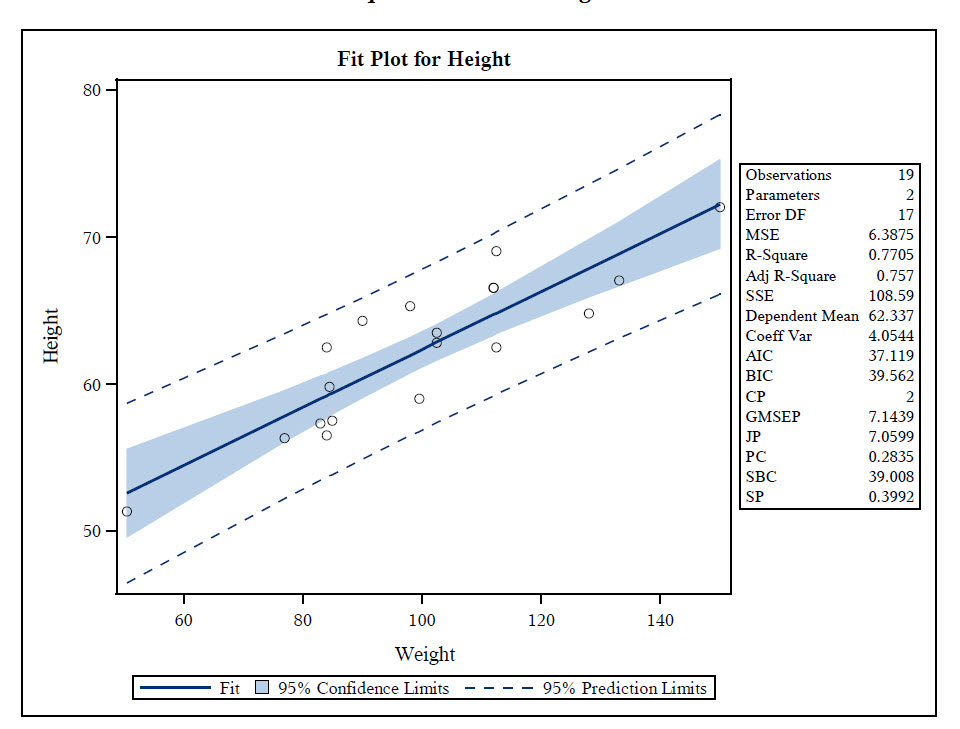
It will likely be removed at the end of the year 2001. We are deprecating residPlot() from the next version of FSA (v0.9.0).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
